Quick start
In this section we will explain how to launch your first job once PhysioFit has been installed onto your system.
See also
If you have already used PhysioFit and are looking for a more in-depth tutorial, check out the Tutorial section.
Graphical user interface
To open the Graphical User Interface, type in a terminal (e.g. Anaconda Prompt if installed on Windows):
physiofit
If you have installed the package in a specific environment, make sure to activate this environment before starting PhysioFit.
PhysioFit interface will open in a new browser window:
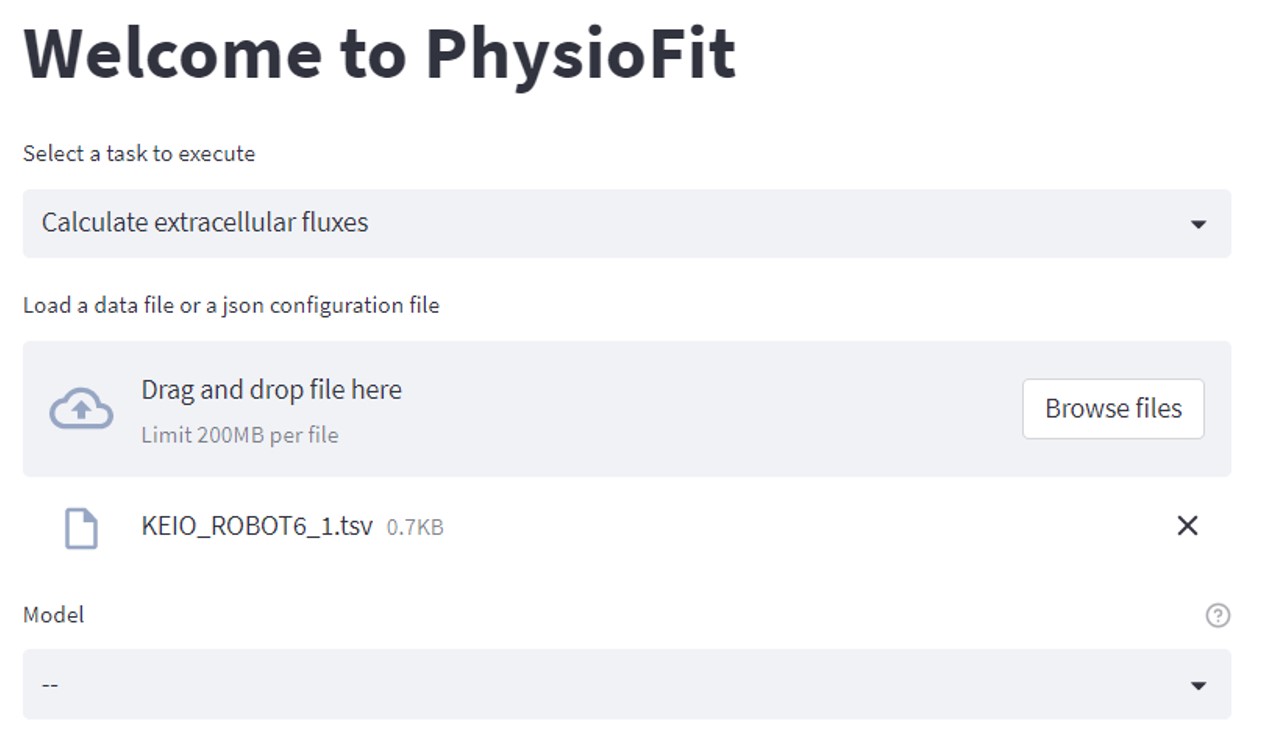
Select an input file (which can be a tsv file containing the data or a yaml configuration file containing the run
parameters and a path towards the data, see Tutorial for more details), select a model, modify the calculation parameters according
to your data, and click on Run flux calculation. PhysioFit proceeds automatically to the flux calculation
and display its progress and possibly important messages such as errors. The output of the calculations (i.e. fluxes and associated statistics)
will be written in a text file as will the statistical test results, while plots will be generated individually for each metabolite (svg files) and combined in a
multi-page pdf file. If multiple experiments were included in the input data, a summary (csv file)
will also be generated. See Output files for more details.
Command line interface
To process your data, type in a terminal:
physiofit [command line options]
Here after the available options with their full names are enumerated and detailed.
usage: physiofit [-h] [-d DATA] [-c CONFIG] [-m MODEL] [-g] [--list] [-v]
[-oc OUTPUT_CONFIG] [-or OUTPUT_RECAP] [-oz OUTPUT_ZIP]
Named Arguments
- -d, --data
Path to data file in tabulated format (txt or tsv)
- -c, --config
Path to config file in yaml format
- -m, --model
Which model should be chosen. Useful only if generating related config file
- -g, --galaxy
Is the CLI being used on the galaxy platform
Default:
False- --list
Return the list of models in model folder
Default:
False- -v, --debug_mode
Activate the debug logs
Default:
False- -oc, --output_config
Path to output the yaml config file
- -or, --output_recap
Path to output the summary
- -oz, --output_zip
Path to export zip file
Online use (W4M)
PhysioFit is available for use online on the Workflow4Metabolomics Galaxy instance (W4M). The tool can be found in the “Isotopic Studies” section. PhysioFit on W4M leverages the configuration file feature to allow users to run the tool without having to input parameters through the Graphical User Interface. To run the tool, load the dataset and the configuration file, and click on “Run”. It’s as simple as that.
W4M also enables the creation of automated workflows, allowing users to build their own data processing pipelines. PhysioFit was designed to make this process as straightforward as possible, by limiting interaction with the GUI and by using the configuration file to set up run parameters. For more information on using and building workflows on the W4M platform, please have a look to the Galaxy training network.
Library
PhysioFit is also available as a library (a Python module) that you can import directly in your Python scripts:
import physiofit
See also
Have a look at our API if you are interested in this feature.